Better
together!
Deciphering microbial gene cluster function using machine learning 🔍
🧪 What is this project about?
In bacterial genomes, genome organization is indicative of function, and proteins that can be functionally grouped into the same pathway tend to be encoded as genes that are in close proximity to each other in the genome. To put it simply: genes that work together are often grouped together as gene clusters! 🧬 In this project, we're developing machine learning (ML) methods, which can be used to discover novel gene clusters in bacterial (meta)genomes.
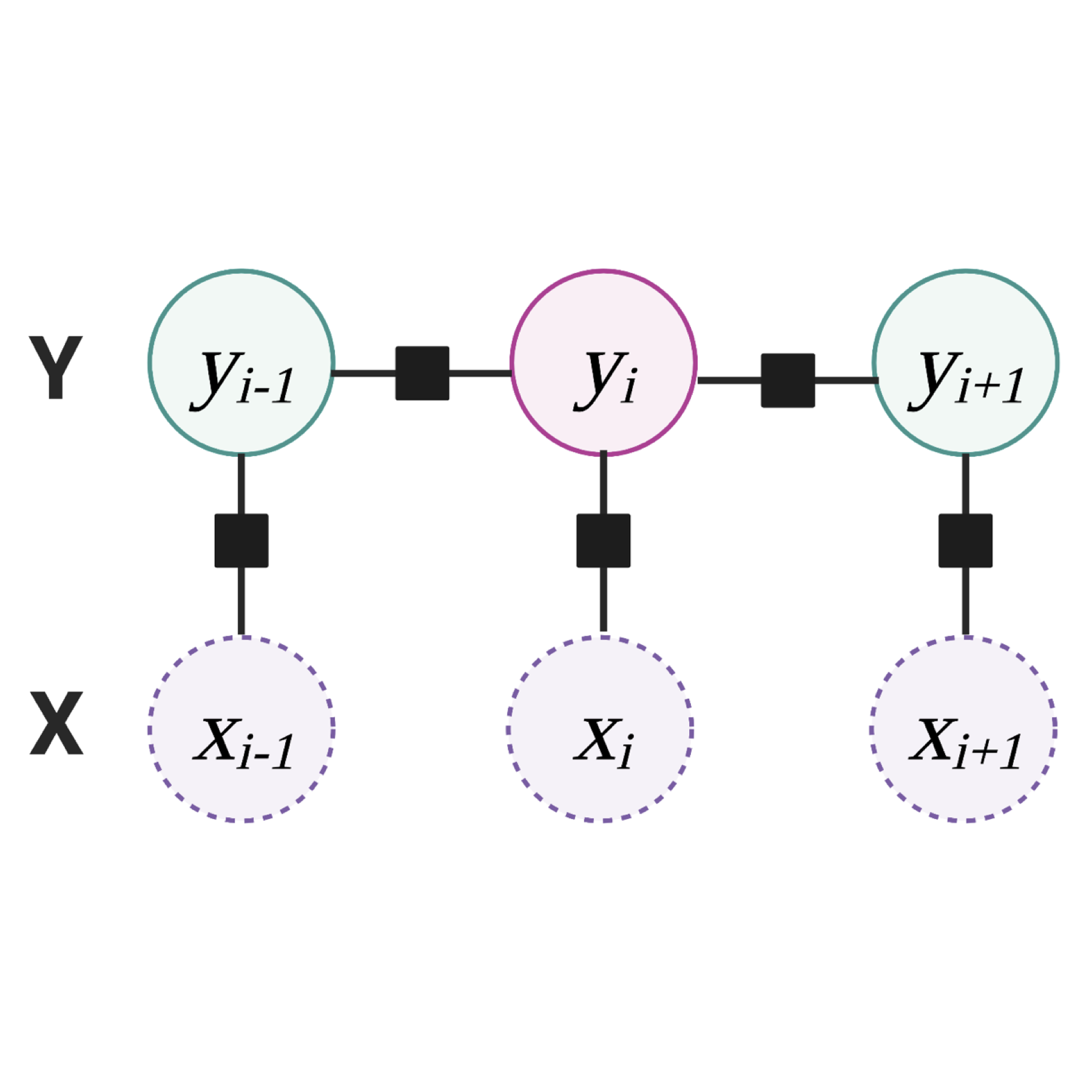
🧐 Why is it important to research this?
Some of the most fascinating and important functions that bacteria can perform are the result of gene clusters...basically, many gene clusters can equip their bacterial hosts with superpowers 🏆, including the ability to produce novel antibiotics or drugs (e.g., biosynthetic gene clusters); the ability to cause illness (e.g., secretion systems; virulence factor-harboring prophage); the ability to resist antibiotics (e.g., antibiotic resistance gene-harboring prophage); and much much more!
🤞 What can we hope to get out of this project?
The methods and tools developed through this project will allow us to discover novel bacterial gene clusters with important roles in human health.