Welcome to the resistance
Phylodynamic methods for antimicrobial resistance surveillance 💊
🧪 What is this project about?
Antibiotics and antimicrobial drugs are used to treat infections caused by dangerous pathogenic bacteria. However, sometimes, these drugs lose their effectiveness...and that's when things get scary, because infections caused by antimicrobial resistant (AMR) bacteria are challenging--and sometimes, impossible--to treat. This project aims to improve our understanding of antimicrobial resistant bacteria by figuring out where they are, which drugs they are resistant to, and why they are resistant to those drugs--using tree-like structures called phylogenies!
🧐 Why is it important to research this?
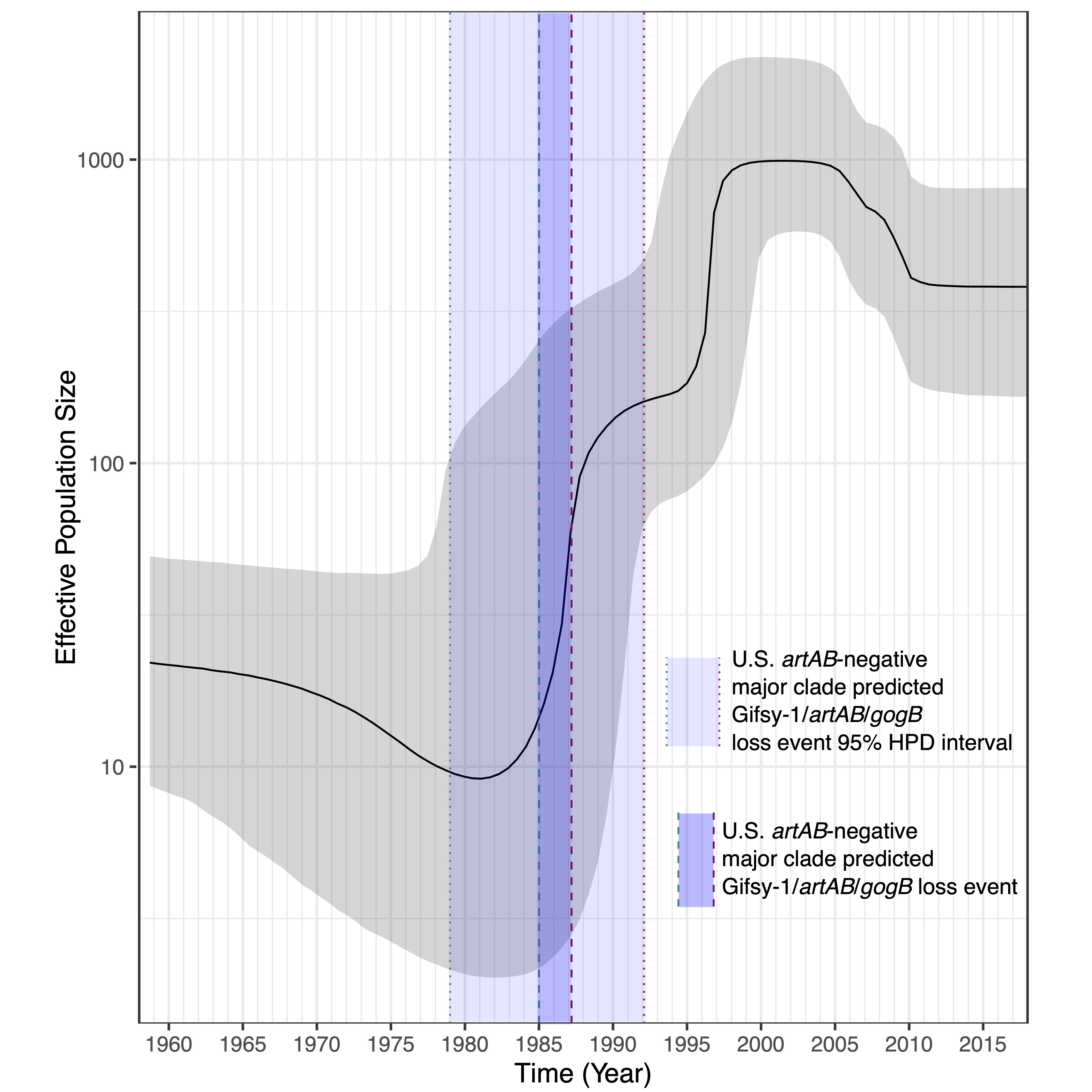
According to the World Health Organization (WHO), AMR is one of the greatest threats to global public health today. WHO and other public health agencies have called for improved monitoring of AMR, and as a result, massive amounts of whole-genome sequencing (WGS) data derived from AMR pathogens are being generated. We're using these massive data sets to model AMR gain, loss, and transmission between hosts, geographic regions, and pathogens!
🤞 What can we hope to get out of this project?
Results obtained from this project can be used to develop targeted intervention strategies, which can be used to mitigate the spread of AMR!